|
Genome_build: Human reference genome (GRCh38. Geneious includes an extensive set of functions to perform sequence alignments of sample reads against reference genomes. HISAT2 (RRID:SCR_015530) was used for read mapping to align processed reads to human reference genome (GRCh38.p13 NIH).įeatureCounts (A part of Subread RRID:SCR_009803) was used to count aligned reads per gene. Geneious Prime is one of many bioinformatics tools that we use to post-process sequencing datasets. Total RNA extraction and preamplification of synthesized cDNAs were performed in Fluidigm C1 systems using the script of 'mRNA Seq HT:RT & Amp (1912x)’.Ī total of 1,600 3'-end enriched scRNA-seq libraires (400 libraries per cell line) were constructed using Nextera XT index Kit v2 Set A/B and Nextera XT DNA library preparation kits.ģ'-end enriched scRNA-seq using Fluidigm C1 systemīasecalling performed using Real-Time Analysis (RTA) version 2.0įluidigm mRNA Sequencing High Throughput Demultiplexer Script ( ) and Geneious Prime 2019.2.1 ( ) were used to demultiplex.īBDuk Trimmer (a part of Bestus Bioinformatics Tools RRID:SCR_016968) was used for quality trimming. It’s particularly good for microbial assemblies with the unique capability to produce circular contigs. Instructions on how to connect to the CAM VPN are. The Geneious Assembler is exible enough to handle data from any type of sequencing machine with reads of any length, including paired-reads and mixtures of reads from different sequencing machines. Individual cells were captured in chambers of Fluidigm HT mRNA-seq IFC. Reliable reference mapping with exclusive mapping algorithms and flexible de novo genome assembly with alignment tools available for comparison and finishing. Download Geneious Prime Note: You must be connected to the CAM VPN if off-site of UCONN Health Center. Upon purchase, cells were stored at -80✬ freezer and then thawed just before directly loading to C1 HT mRNA-seq IFCs. 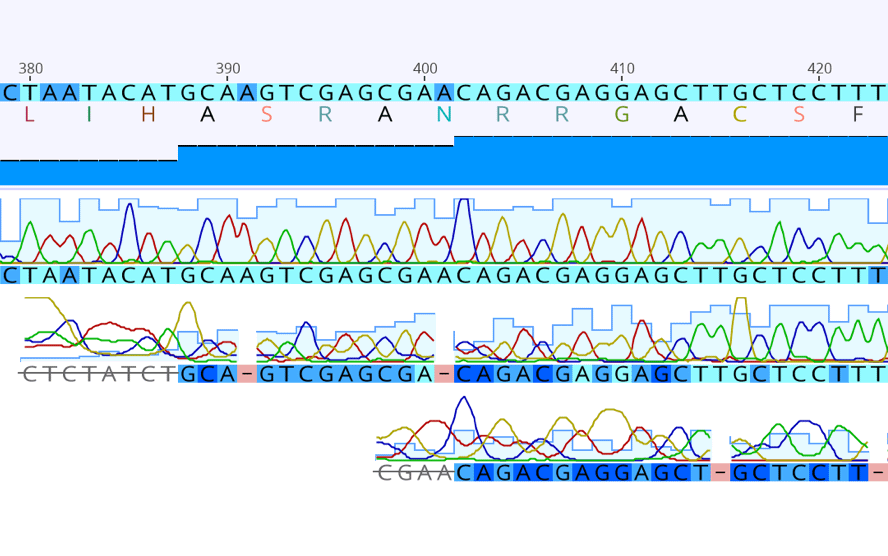
If there is a fasta file with the same name, and in the. When you import the annotation file you will be prompted for the sequence in one of two ways: 1. On Windows and Linux, copy the le to the Geneious installation directory and rename it to default user preferences.xml. To do this, you can either import the sequence from a fasta file at the same time you import the annotation file, or you can import the file onto an existing sequence in your Geneious database. 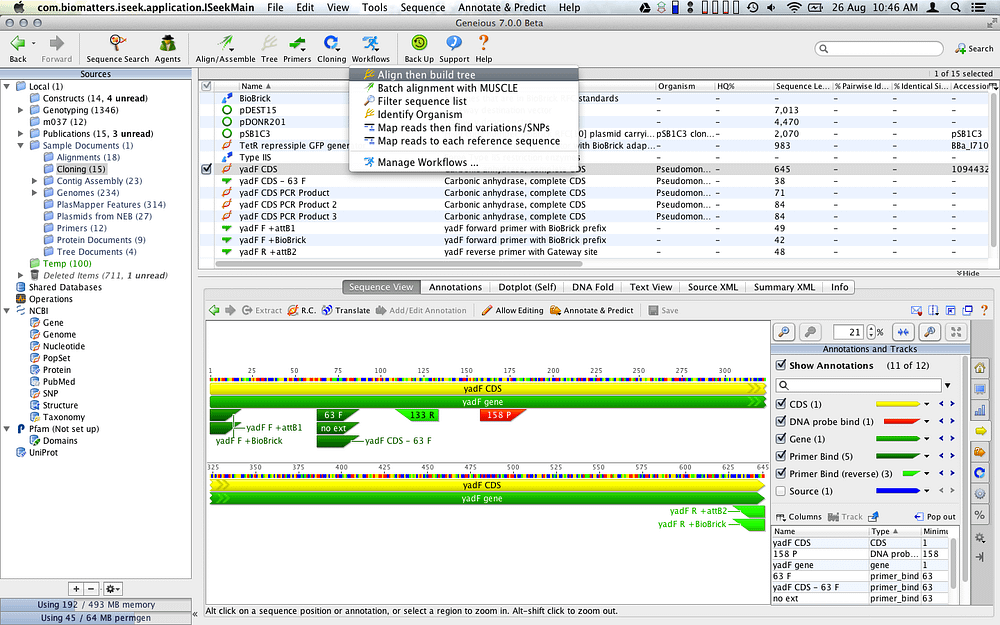
Shut down and then copy the le Geneious 2022.1 Data/user preferences.xml. 
Start a fresh copy of Geneious Prime, set it up the way you want. Geneious is extremely handy and convenient for small scale analyses. Geneious makes it simple to map sequences and easily edit your contigs. Geneious is meant to be an easy to use interface for biologists and not a replacement for a proper bioinformatician. GEO help: Mouse over screen elements for information. 22.2.1 Change preferences within Geneious Prime. Using Geneious Prime for your Sanger sequencing analysis will improve your mapping accuracy, decrease your analysis time and streamline your processes.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
